Conversion to time series format¶
For a lot of applications it is favorable to convert the image based format into a format which is optimized for fast time series retrieval. This is what we often need for e.g. validation studies. This can be done by stacking the images into a netCDF file and choosing the correct chunk sizes or a lot of other methods. We have chosen to do it in the following way:
Store the grid points in a 1D array. This also allows reduction of the data volume by e.g. only saving the points over land.
Store the time series in netCDF4 in the Climate and Forecast convention Orthogonal multidimensional array representation
Store the time series in 5x5 degree cells. This means there will be 2566 cell files and a file called
grid.ncwhich contains the information about which grid point is stored in which file. This allows us to read a whole 5x5 degree area into memory and iterate over the time series quickly.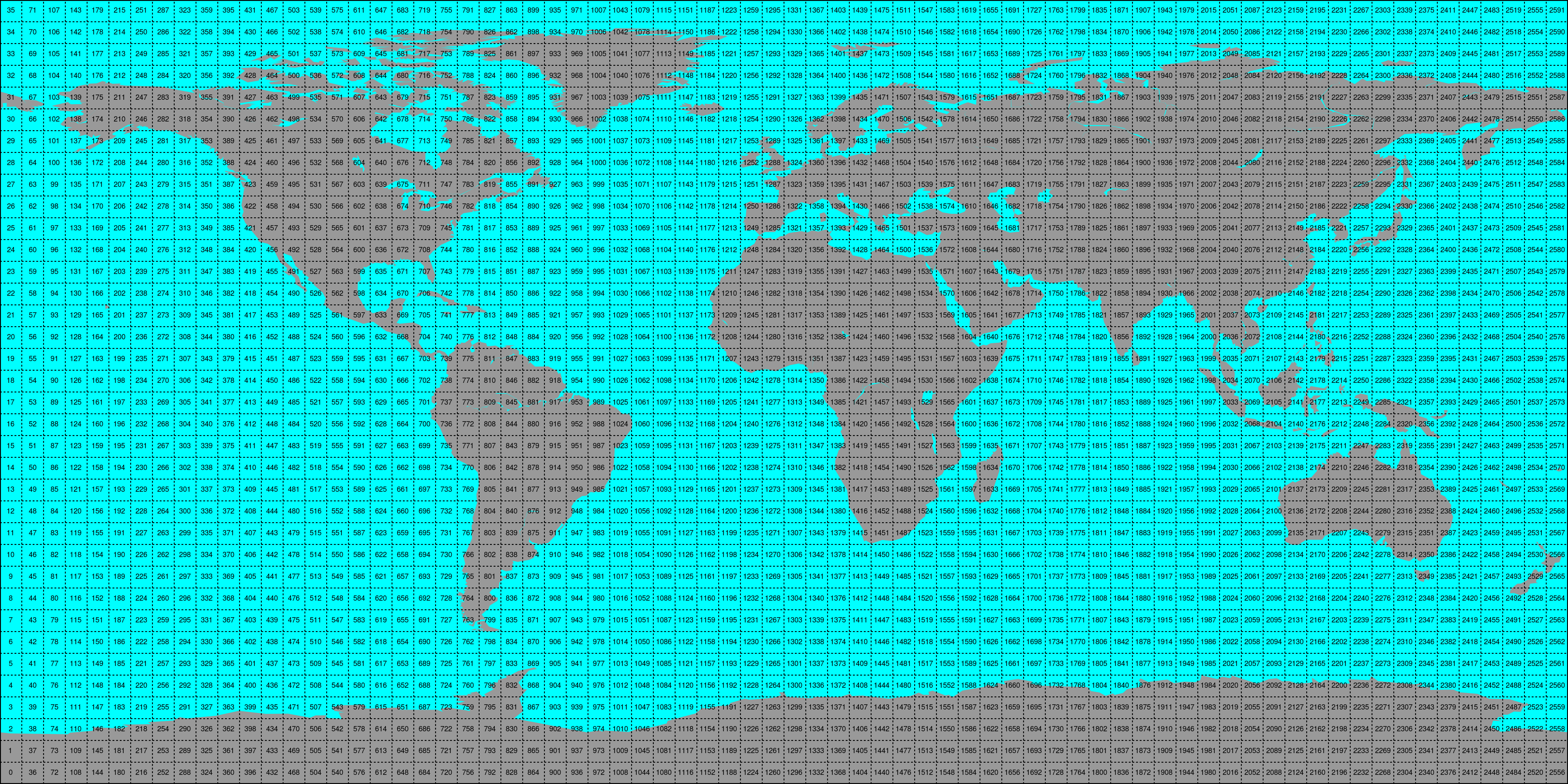
SPL3SMP¶
This conversion can be performed using the smap_repurpose command line
program. An example would be:
smap_repurpose /SPL3SMP_data /timeseries/data 2015-04-01 2015-04-02 soil_moisture soil_moisture_error --overpass AM
Which would take SMAP SPL3SMP data stored in /SPL3SMP_data from April 1st
2015 to April 2nd 2015 and store the parameters soil_moisture and
soil_moisture_error for the AM overpass as time series in the
folder /timeseries/data. When the PM overpass is selected, time series variables
will be renamed with the suffix _pm.
Conversion to time series is performed by the repurpose package in the background. For custom settings
or other options see the repurpose documentation and the code in
smap_io.reshuffle.
Note: If a RuntimeError: NetCDF: Bad chunk sizes. appears during reshuffling, consider downgrading the
netcdf4 library via:
conda install -c conda-forge netcdf4=1.2.2